ShapeFactor¶
[1]:
%pylab inline
Populating the interactive namespace from numpy and matplotlib
[2]:
import logging
import json
import pyhf
from pyhf import Model
logging.basicConfig(level = logging.INFO)
[3]:
def prep_data(sourcedata):
spec = {
'channels': [
{
'name': 'signal',
'samples': [
{
'name': 'signal',
'data': sourcedata['signal']['bindata']['sig'],
'modifiers': [
{
'name': 'mu',
'type': 'normfactor',
'data': None
}
]
},
{
'name': 'bkg1',
'data': sourcedata['signal']['bindata']['bkg1'],
'modifiers': [
{
'name': 'coupled_shapefactor',
'type': 'shapefactor',
'data': None
}
]
}
]
},
{
'name': 'control',
'samples': [
{
'name': 'background',
'data': sourcedata['control']['bindata']['bkg1'],
'modifiers': [
{
'name': 'coupled_shapefactor',
'type': 'shapefactor',
'data': None
}
]
}
]
}
]
}
pdf = Model(spec)
data = []
for c in pdf.spec['channels']:
data += sourcedata[c['name']]['bindata']['data']
data = data + pdf.config.auxdata
return data, pdf
[4]:
source = {
"channels": {
"signal": {
"binning": [2,-0.5,1.5],
"bindata": {
"data": [220.0, 230.0],
"bkg1": [100.0, 70.0],
"sig": [ 20.0, 20.0]
}
},
"control": {
"binning": [2,-0.5,1.5],
"bindata": {
"data": [200.0, 300.0],
"bkg1": [100.0, 100.0]
}
}
}
}
data, pdf = prep_data(source['channels'])
print('data: {}'.format(data))
init_pars = pdf.config.suggested_init()
print('expected data: {}'.format(pdf.expected_data(init_pars)))
par_bounds = pdf.config.suggested_bounds()
INFO:pyhf.pdf:Validating spec against schema: /home/jovyan/pyhf/src/pyhf/data/spec.json
INFO:pyhf.pdf:adding modifier mu (1 new nuisance parameters)
INFO:pyhf.pdf:adding modifier coupled_shapefactor (2 new nuisance parameters)
data: [220.0, 230.0, 200.0, 300.0]
expected data: [120. 90. 100. 100.]
[5]:
print('initialization parameters: {}'.format(pdf.config.suggested_init()))
unconpars = pyhf.optimizer.unconstrained_bestfit(pyhf.utils.loglambdav, data, pdf,
pdf.config.suggested_init(), pdf.config.suggested_bounds())
print('parameters post unconstrained fit: {}'.format(unconpars))
initialization parameters: [1.0, 1.0, 1.0]
parameters post unconstrained fit: [0.99981412 2.00002042 3.00006469]
/home/jovyan/pyhf/src/pyhf/tensor/numpy_backend.py:173: RuntimeWarning: divide by zero encountered in log
return n * np.log(lam) - lam - gammaln(n + 1.0)
[6]:
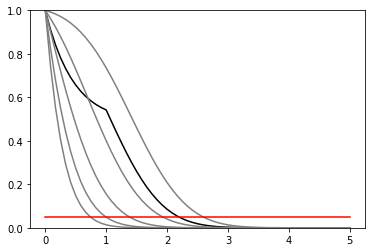
def plot_results(testmus, cls_obs, cls_exp, poi_tests, test_size = 0.05):
plt.plot(poi_tests,cls_obs, c = 'k')
for i,c in zip(range(5),['grey','grey','grey','grey','grey']):
plt.plot(poi_tests, cls_exp[i], c = c)
plt.plot(testmus,[test_size]*len(testmus), c = 'r')
plt.ylim(0,1)
def invert_interval(test_mus, cls_obs, cls_exp, test_size=0.05):
crossing_test_stats = {'exp': [], 'obs': None}
for cls_exp_sigma in cls_exp:
crossing_test_stats['exp'].append(
np.interp(
test_size, list(reversed(cls_exp_sigma)), list(reversed(test_mus))
)
)
crossing_test_stats['obs'] = np.interp(
test_size, list(reversed(cls_obs)), list(reversed(test_mus))
)
return crossing_test_stats
poi_tests = np.linspace(0, 5, 61)
tests = [pyhf.utils.hypotest(poi_test, data, pdf, init_pars, par_bounds, return_expected_set=True)
for poi_test in poi_tests]
cls_obs = np.array([test[0] for test in tests]).flatten()
cls_exp = [np.array([test[1][i] for test in tests]).flatten() for i in range(5)]
print('\n')
plot_results(poi_tests, cls_obs, cls_exp, poi_tests)
invert_interval(poi_tests, cls_obs, cls_exp)
[6]:
{'exp': [0.741381412468345,
0.9949353526744877,
1.3845144105754894,
1.9289946435921614,
2.594077794516857],
'obs': 2.1945970333027187}